Spatial transcriptomic analysis of pediatric lupus nephritis at single cell resolution
View On Demand
Abstract
Despite recent FDA approval of new therapies to treat lupus nephritis, most patients fail to achieve clinical remission with standard-of-care treatment. This emphasizes the need for a greater understanding of the pathophysiology of lupus nephritis, with the expectation that this knowledge will inform the development of effective, targeted therapies for lupus nephritis.
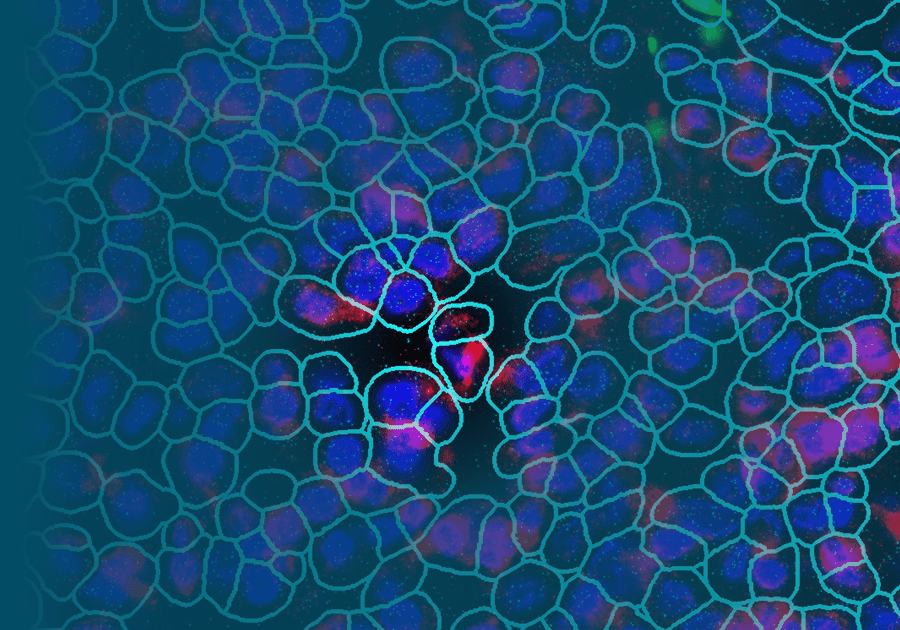
To date, pre-clinical studies have typically used animal models to identify immune pathways driving disease, but these efforts have been plagued by limited correlation with clinical benefit in humans. For this reason, the ability to directly analyze human kidney tissue from lupus nephritis patients is of significant importance. We performed spatial transcriptomics of pediatric Class III and Class IV lupus nephritis using the CosMx Spatial Molecular Imager (Nanostring, Inc.). A significant advantage of the CosMx platform is the ability to study archived clinical kidney biopsy tissue stored in FFPE. In the pilot phase of our study, we confirmed the accurate annotation of major kidney stromal and immune cell types, including rare cells such as podocytes that have been challenging to identify in single-cell RNA-Seq datasets. Our ongoing analyses are focused on the spatial location of immune cells infiltrating the lupus nephritis kidney, the stromal signatures of inflammation-mediated injury, and whether gene expression differs between affected and unaffected glomeruli across tissue samples.
In addition to advancing our understanding of lupus nephritis pathophysiology, we anticipate that these efforts will generate a spatially-resolved cell atlas of human lupus nephritis that will be an important resource for investigators in academia and industry.
Speaker



