nCounter® Viral Panel Plus Menu
Helping Your Research
There are many applications in infectious disease research where viral detection is useful, such as understanding how the viral life cycle affects the host response, disease pathogenesis, the impact of antivirals and vaccines, and chronic infection. It can be difficult with RNA sequencing to detect viral transcripts in infected samples due to the overwhelming amount of host material present.

How it Works
The targeted and customizable nature of nCounter technology makes it easy to profile transcripts from the host and one or more pathogens simultaneously. Probes for pathogens can be added as a Panel Plus spike-in to an off-the-shelf human or mouse Gene Expression Panel or included in a de novo Custom CodeSet. The nCounter Viral Panel Plus Menu is a list of pre-designed Research Use Only probes for eight viruses associated with chronic disease and/or cancer. You can use the pre-designed probes as is in a Panel Plus design or mix and match probes from different viruses to create your own Panel Plus of choice. Probes were designed to cover as many sequences as possible while at the same time ensuring sensitivity, specificity, and breadth of coverage.
Probes cannot be used in a standalone assay and must be paired with an nCounter Gene Expression Panel. Check with bioinformatics about compatibility with a specific panel; however, the Viral Panel Plus Menu has been designed to be compatible with most nCounter panels. Probes are also compatible with Elements™ chemistry and PlexSet™ assays. Viral detection may differ across different sample types.


For ideal coverage of the host response to a particular virus, pair the Viral Panel Plus Menu probes with the Host Response Panel or study the effect of chronic viral infections on T cell, B cell, and NK cell function with the Immune Exhaustion Panel.

Panel Selection Tool
Find the gene expression panel for your research with easy to use panel pro
Find Your Panel
Publications
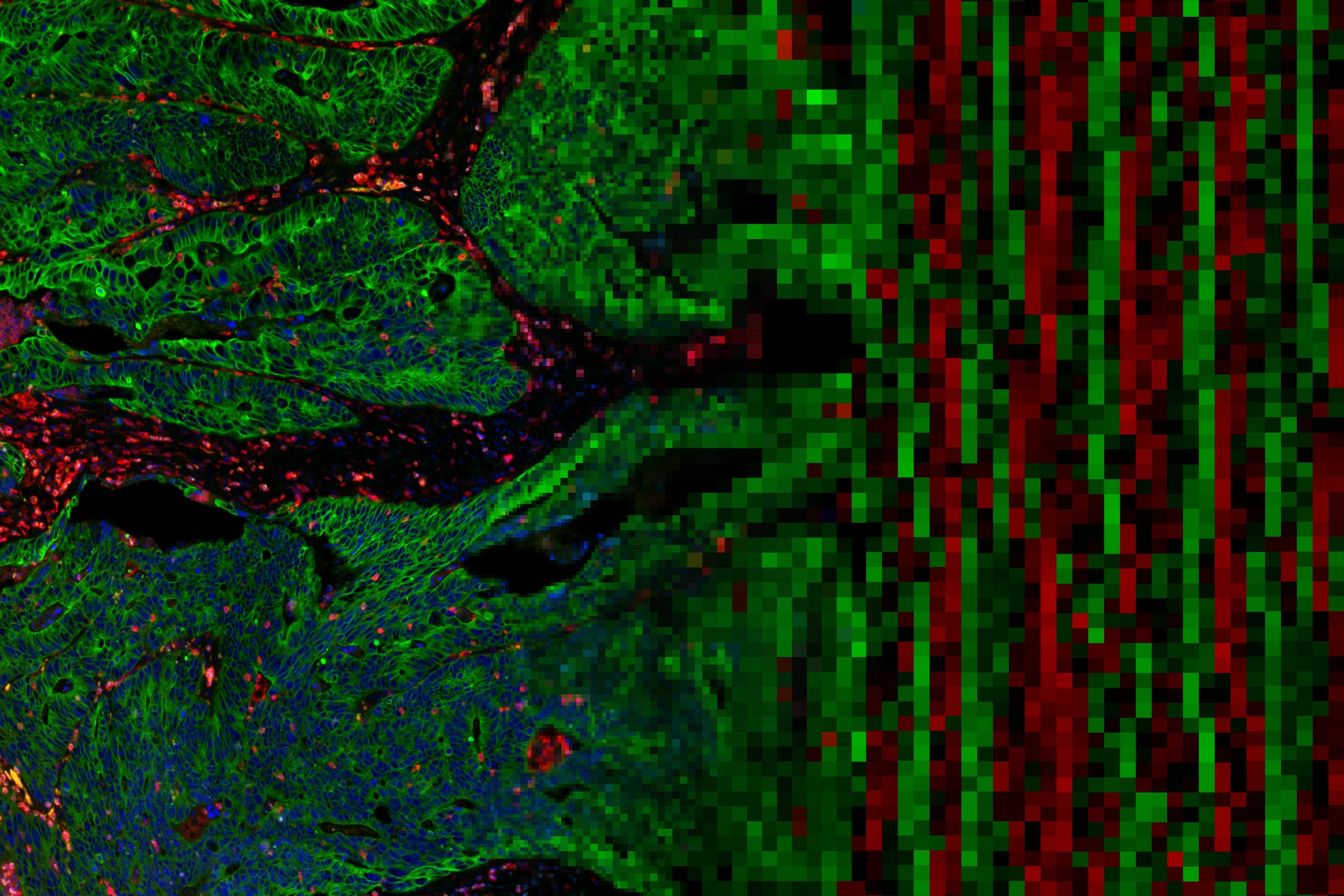
Single-cell and spatial multi-omics highlight effects of anti-integrin therapy across cellular compartments in ulcerative colitis
Ulcerative colitis (UC) is driven by immune and stromal subsets, culminating in epithelial injury. Vedolizumab (VDZ) is an anti-integrin antibody that is effective for treating UC.
In situ single-cell profiling sheds light on IFI27 localisation during SARS-CoV-2 infection
The utilization of single-cell resolved spatial transcriptomics to delineate immune responses during SARS-CoV-2 infection was able to identify M1 macrophages to have elevated expression of IFI27 in areas of infection.
The SARS-CoV-2 pandemic has affected over 600 million people to date, resulting in over 6.
Whole transcriptome profiling of placental pathobiology in SARS‐CoV‐2 pregnancies identifies placental dysfunction signatures
Objectives: Severe Acute Respiratory Syndrome Coronavirus 2 (SARS‐CoV‐2) virus infection in pregnancy is associated with higher incidence of placental dysfunction, referred to by a few studies as a ‘preeclampsia‐like syndrome’. However, the mechanisms underpinning SARS‐CoV‐2‐induced placental malfunction are still unclear.
Support Documents

Contact Us
Have questions or simply want to learn more?
Contact our helpful experts and we’ll be in touch soon.