
The Votes Are In- What Causes qPCR and RNA-Seq Variability?
Retraction. A word feared by every scientist conducting lab experiments and publishing data. I came across an interesting article recently entitled “Sources of error in the retracted scientific literature” (Arturo Casadevall et. al.). The paper noted that retraction of flawed articles is an important mechanism for correction of scientific literature and analyzed 423 retraction notices for articles indexed in PubMed for which no misconduct was identified. They found the most common causes of error-related retraction to be lab errors, analytical errors, and irreproducible results. Now, most researchers I know are inherently good people, so I wanted to further explore where variability could be coming from in some common research platforms.
At the American Society for Human Genetics annual meeting (ASHG) this year, NanoString® conducted a poll open to conference attendees that elected to participate. The poll asked a simple question:
What is the biggest source of variability in qPCR or RNA-Seq workflows?
Participants could select from the following options, and no limitations were placed on the number of selections made:
- Bioinformatics/Data Analysis
- Reverse Transcription
- Amplification
- Pipetting/Human Factors
- Normalization
- RNA extraction/ Sample prep
Over three days, 116 votes were collected, and here are the ultimate findings:
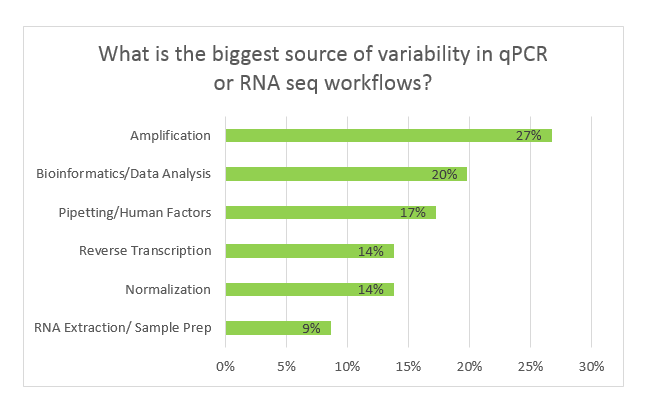
Far and wide, most researchers believe the process of creating cDNA and amplifying the target introduces the most variability into the workflow. These are very logical selections, as Reverse Transcription (RT) is known to introduce errors into transcripts and amplification bias is a concept universally recognized.
Bioinformatics/data analysis was another highly rated factor impacting variability. Several poll participants commented on qPCR assay design or the method chosen to analyze sequencing data highly impacting results. Similarly, normalization and the choice of reference gene selection is also known to have a significant impact on obtained results. I recall reading a paper “Critical appraisal of quantitative PCR results in colorectal cancer research: Can we rely on published qPCR results?” (JR Dijkstra et. al) where the authors assessed 179 colorectal cancer publications that made use of RT-qPCR from 2006- August 2013 for the number of reference genes used and whether they had been validated. The validity of only 3% of the publications could be adequately assessed! Clearly even with MIQE guidelines there is room for vast improvement in normalization and the analysis front.
Pipetting/human factors was an option selected by 17% of participants. Several participants commented that automated workflows alleviate this factor from having high impact. Some brave souls, however, admitted that yeah- they mess up an awful lot. Kudos to you brave souls for taking one for the team in the name of research.
The final and least popular selection was RNA extraction/sample prep. Only 9% of participants selected this bucket, and often the individuals were employees of genomic tools provider companies.
The way to overcome these variabilities? Through an RNA direct-detection based platform like NanoString’s nCounter® Technology. Check out this video to learn more.
So, there you have it folks! A very interesting and insightful look into the causes of variability through the eyes of researchers. If you would like more information on how NanoString’s direct digital detection technology overcomes these variability barriers, please visit our Challenges of RT resource page.
FOR RESEARCH USE ONLY. Not for use in diagnostic procedures.

