Immunotherapy in Tumors: Integrated Information to Outsmart Cancer
If it wasn’t such a devastating disease, there would be memes about how super-smart cancer is.
Cancer cells can express neoantigens — tumor-specific, mutated peptides — that, when processed and presented by the MHC to T cells, mount an adequate anti-tumor response. Yet, cancer cells can also sneakily bypass elimination by manipulating the tumor microenvironment to be more immunotolerant, exploiting natural immune checkpoints such as the Programmed Death 1 (PD-1)/Programmed Death Ligand-1 (PD-L1) and the CTLA-4/B7 immunosuppressive pathways.
A recent generation of drugs knowns as “Immune Checkpoint Inhibitors” (ICIs) have revolutionized cancer treatment by targeting these pathways used by cancer cells to evade the immune system. These immunotherapies have been shown to elicit an anti-tumor response in multiple cancers, but the response varies across different patients with the same type of cancer.
Responses to ICIs can occur preferentially in tumors wherein there is a pre-existing adaptive immune response, a so-called ‘T cell inflamed’ phenotype. This has typically been measured by immunohistochemistry (IHC) by analysis of PD-L1 expression and/or by gene expression profiling for a T Cell inflamed gene signature, in the hope that classifying tumors based on whether they are ‘inflamed’ or ‘hot’ may help predict response to checkpoint inhibitors.
However, an inflamed tumor alone does not necessarily guarantee response to ICI therapy. In fact, response to these therapies has been shown to be associated with tumors that have extensive antigenicity, typically quantified as tumor mutational burden (TMB), microsatellite instability (MSI), or Mismatch Repair (MMR) deficiency: mechanistically, MSI-High tumors are actually a type of tumor characterized by high TMB, wherein a deficient mismatch repair system generates a high mutational load. Tumor antigenicity is generally tested by NGSs and/or IHC.
It is rapidly becoming clear that no one single measurement of immune phenotype and/or tumor antigenicity can be expected to fully capture the potential outcome of treatment with an ICI. Stratifying tumors holistically based on their T cell inflamed phenotype, TMB, MSI, and/or MMR is desired, but this requires using multiple disparate technologies, which is not ideal. Understandably, a major hurdle for immuno-oncology (IO) research is to get as much integrated and actionable biological information as possible in a short period of time from a small amount of tumor tissue.
NanoString has accepted this challenge and raised it to accelerate IO research with the nCounter® System and its portfolio of ready-to-use gene expression (GX) panels such as the PanCancer IO360™ Panel.
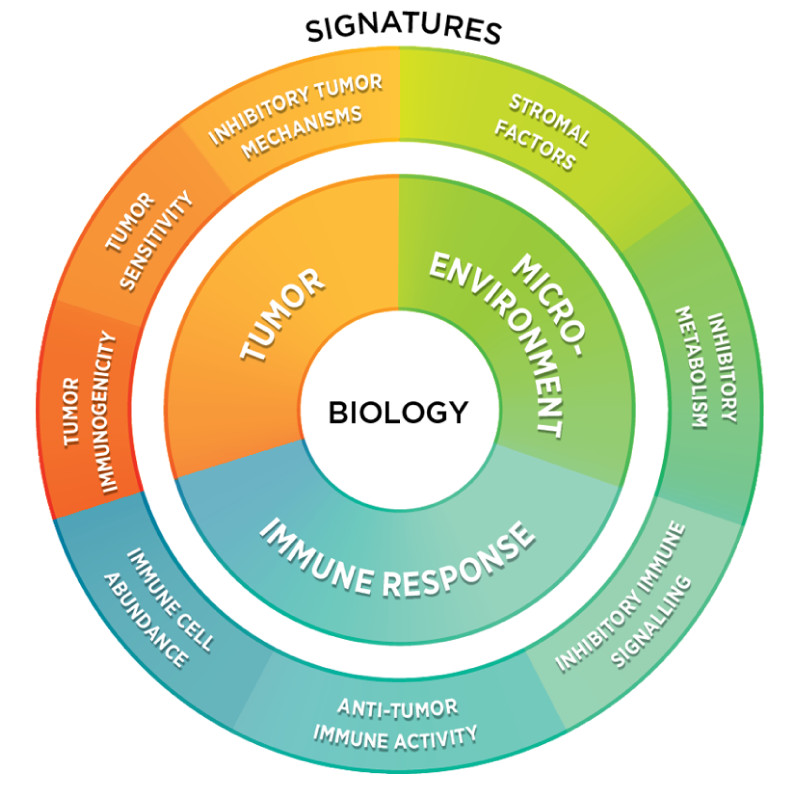
Designed for Research Use Only (RUO), the IO 360 Panel and its curated content was developed in conjunction with leading IO researchers across academia and industry. The panel enables digital detection and differential expression profiling of 770 genes involved in the interplay between the tumor, microenvironment, and immune response. The panel includes 48 biological signatures such as a tumor immunogenicity signature for the loss of MMR genes and the Tumor Inflammation Signature (TIS), an 18-gene signature that measures the extent of a pre-existing adaptive immune response that may be associated with the response of ICIs. As with all NanoString GX panels, data is obtained within 24 hours with minimal hands-on time and as little as one FFPE tissue section: the assay is robust, specific and works well with degraded RNA.

The IO 360 Panel comes with the option of an easy-to-interpret and interactive Data Analysis Report with scores for all the 48 signatures. The report is generated and delivered by a team of NanoString scientists and supporting biostatisticians and is comprised of an executive summary, statistical methods section, and publication-ready and annotated graphs and figures
Combined with the power of the Data Analysis Report and the 48 signature scores, the IO 360 Panel allows researchers to integrate differential expression data on intrinsic DNA repair pathways with that of the immune activation state within the tumor and the microenvironment, moving closer to a more holistic picture of the biology of a given tumor and better stratification for ICI response.
For Research Use Only. Not for use in Diagnostic Procedures.
References:
Danaher P et al., Journal for Immuno Therapy of Cancer (2019) 7:15 DOI: 10.1186/s40425-018-0472-1
Dung et al., Science. 2017 Jul 28; 357(6349): 409–413. DOI: 10.1126/science.aan6733
Ton N. Schumacher, Robert D. Schreiber, Science. 03 Apr 2015: Vol. 348, Issue 6230, pp. 69-74 DOI: 10.1126/science.aaa4971


