COVID-19 Research Solutions
Helping Your Research
NanoString, the leader in Spatial Biology and Spatial Transcriptomics, is committed to advancing scientific understanding of the impact of the coronavirus SARS-CoV-2 on human health worldwide. As many of our customers shift their work to addressing urgent needs around therapeutic & vaccine development, and studying viral pathogenesis, we are joining them in this effort.
NanoString offers a broad range of high-plex gene expression panels enabling research on pathogenesis and the host response. In several recently published studies, nCounter® gene expression assays such as the Immunology Panel and Inflammation Panel were used to describe the dynamic and varied host response that shapes the progression of COVID-19. Building on this success, NanoString developed the 785-plex Human and Mouse Host Response Panel as well as a Coronavirus Panel Plus spike-in, enabling measurement of the SARS-CoV-2 virus, additional human coronaviruses, and the human ACE2 receptor RNA in addition to the measurement of host immune response genes. The Coronavirus Panel Plus can be spiked-in to any gene expression panel, expanding the range of biology that can be studied with COVID-19 samples.
- The GeoMx® Digital Spatial Profiler (DSP) enables simultaneous high-plex spatial analysis of pathogens and the host response with FFPE and fresh frozen tissue.
- The GeoMx RNA Barcoding Service and Protein Barcoding Service allows you to spike-in probes for SARS-CoV-2 into standalone GeoMx RNA and Protein Assays.
- The COVID-19 GeoMx-formatted Antibody Panel, developed in partnership with Abcam, is a five-antibody custom protein panel that can be run alongside GeoMx protein assays. These assays include SARS-CoV-2 viral markers and the ACE2 receptor, among other receptors, proteases, cell markers, and viral response markers, and are available through the GeoMx Technology Access Program.
How it Works
nCounter Host Response Panel
The 785-plex nCounter Host Response Panel contains probes for human or mouse genes involved in the host response to pathogens. Best suited for use with blood samples, the Host Response Panel covers the five stages of infection: incubation, prodromal period, peak illness, decline, and convalescence, with probes for genes involved in the innate and adaptive immune response, interferon signaling, host susceptibility, and homeostasis. The panel includes NanoString’s unique immune cell typing signatures for quantifying the relative abundance of 14 different immune cell types, and data from the panel can be easily analyzed in minutes with the nSolver™ Data Analysis Software or the cloud-based ROSALIND™ by OnRamp Bio. The panel is ideally suited for use to study SARS-CoV-2 but can be easily used to study other types of pathogens.
COVID-19 Tissue Reference Gene List
Study organ damage wrought by COVID-19 by adding up to 55 genes to the Host Response Panel as a Panel Plus or by adding custom targets to a GeoMx Digital Spatial Profiling RNA assay. Download the COVID-19 Tissue Reference Gene List to get ideas on which genes to add for studying the effect of COVID-19 on the kidney, GI tract, heart, liver, endothelium, lung, and brain. Check back often for updates as the NanoString Bioinformatics team continually adds to and refines these reference gene lists.
The content is mined from publications and represents genes associated multiple times in the literature to tissue damage in the context of COVID-19, as well as the top ten genes associated with a particular tissue. The rationale for each gene’s inclusion and its presence or absence in the Host Response Panel and the GeoMx COVID-19 Immune Response Atlas is noted. Listed genes can be used alongside any nCounter gene expression panel or GeoMx RNA assay; please contact bioinformatics for more information on which genes may already be present in a given panel or GeoMx RNA assay of interest.
Contact us for more information on COVID-19 research tools from NanoString.
Coronavirus Panel Plus
NanoString has developed a Coronavirus Panel Plus product that can be added as a spike-in to nCounter Gene Expression panels and/or Custom CodeSets
- The Coronavirus Panel Plus is for experimental Research Use Only (RUO)
- Probes are compatible for use with most nCounter Gene Expression Panels
- The Coronavirus Panel Plus cannot be used as an individual assay (as with all Panel Plus products)
Download the Coronavirus Panel Plus Gene List
Probes for SARS-CoV-2 are compatible with Elements and PlexSet™ Reagents.
Important Update: includes probe coverage for all current variants of concern (VOC) as defined by WHO (as of Oct 2022.)
GeoMx DSP COVID-19 Protein and RNA Analysis
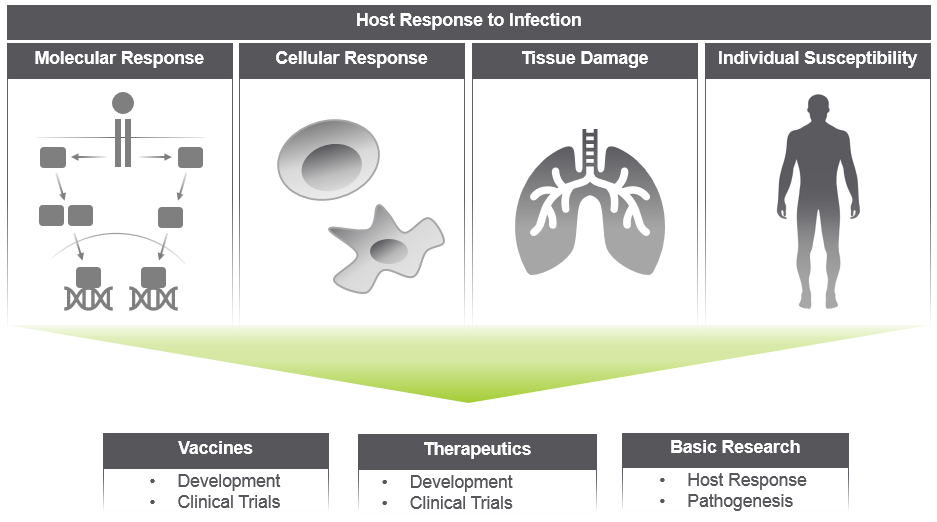
Rapidly perform high-plex spatial analyses of the host response in FFPE or fresh frozen tissue using the GeoMx Digital Spatial Profiler (DSP). NanoString’s GeoMx DSP platform enables high-plex protein and RNA experiments in key areas of biology such as molecular response, cellular (immune) response, tissue damage, and drivers of individual susceptibility to severe forms of disease.
Custom RNA probes for SARS-CoV-2 and related genes can be spiked into the GeoMx Human Whole Transcriptome Atlas to study the pathogenesis and progression of COVID-19 in human samples. RNA targets are profiled simultaneously using the GeoMx DSP and an Illumina next generation sequencer (NGS) for readout. Users can run ADC RNAScope probes alongside GeoMx RNA probes to identify regions of interest.
A five-antibody custom, ready-to-go protein panel, with receptor, protease, and viral markers is available through the GeoMx Technology Access Program or for order through Abcam. This COVID-19 GeoMx-formatted Antibody Panel is run with the 20-plex GeoMx Immune Cell Profiling Core (plus controls) with readout on the nCounter Analysis System. Users can add up to six 10-plex modules including the Immune Activation Status, Immune Cell Typing, and/or Cell Death modules to more deeply profile proteins involved in T cell activation and cell death. NanoString scientists can recommend commercially-available markers for lung epithelium, nasal epithelium, immune response markers, and the viral spike protein.

Panel Selection Tool
Find the gene expression panel for your research with easy to use panel pro
Find Your Panel
Publications
SARS-CoV-2 infection induces a long-lived pro-inflammatory transcriptional profile
Background: The immune response to severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in COVID-19 patients has been extensively investigated. However, much less is known about the long-term effects of infection in patients and how it could affect the immune system and its capacity to respond to future perturbations.
Prevalence and Mechanisms of Mucus Accumulation in COVID-19 Lung Disease
Rationale: The incidence and sites of mucus accumulation, and molecular regulation of mucin gene expression, in COVID-19 lung disease have not been reported. Objectives: Characterize incidence of mucus accumulation and the mechanisms mediating mucin hypersecretion in COVID-19 lung disease.
SARS-CoV-2 infection produces chronic pulmonary epithelial and immune cell dysfunction with fibrosis in mice
A subset of individuals who recover from coronavirus disease 2019 (COVID-19) develop post-acute sequelae of SARS-CoV-2 (PASC), but the mechanistic basis of PASC-associated lung abnormalities suffers from a lack of longitudinal tissue samples. The mouse-adapted severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) strain MA10 produces an acute respiratory distress syndrome (ARDS) in mice similar to humans.
Support Documents
*pre-amplification primers for low input experiments will also be made available.

Want to know more?
Contact us
For more information on COVID-19 research tools from NanoString