Common questions in molecular biology: How is DNA barcoding used in research?
The term DNA barcoding is used in two ways in research. The original use was based on the discovery of a short, standardized genetic region found in fish to have a unique sequence in each species of fish (Hebert). The idea of a unique sequence used for identification of species was conceptually similar to the Universal Product Codes (UPCs) used by supermarkets to identify their items. Where the UPC system uses a unique sequence of different sized bars to identify each item, the system of DNA barcoding uses naturally occurring unique sequences of DNA to identify each species (Hebert). This concept of using unique DNA sequences for barcoding purposes has since been adopted by biologists for a variety of uses, including gene expression studies.
Naturally-occurring DNA barcode libraries
The discovery of short, standardized genetic regions containing sequences specific to each species led to the development of a library of sequences for use in identifying individual specimens. The original gene region identified was the mitochondrial cytochrome C oxidase 1 (COI) subunit gene. To identify a new specimen, a 648-bp region of the COI gene is sequenced after amplification by PCR. The DNA sequence is analyzed against libraries of reference sequences (Ratnasingham and Hebert, 2007). Barcoding of this type was initiated in 2004, followed by development of the International Barcode of Life project in 2010 (International Barcode of Life 2010).
DNA barcoding was dubbed “the renaissance of taxonomy” (Miller 2007) since it engendered better species-identification and increased information about biodiversity. This use of DNA barcoding was thought to have the potential to “accelerate our discovery of new species, improve the quality of taxonomic information and make this information readily available to nontaxonomists and researchers outside of major collection centers” (Miller 2007). DNA barcoding continues to be used in species identification, with the reference library of species sequences growing exponentially.
Engineered DNA barcodes
Inspired by the idea of using unique DNA sequences for identification purposes, scientists began using engineered sequences in a variety of ways. Initial DNA barcodes were used in a manner similar to that for identifying species, using PCR and sequencing to identify signature-tagged mutants (Mazurkiewicz), sequences from multiplexed high-throughput pyrosequencing (Parameswaran), and insertions in viral vectors (Chen). In these methods, a known unique sequence is inserted into an engineered DNA construct, such as a mutant library, for recovery after a functional screen to confirm the presence of the mutant construct (Mazurkiewicz).
Initial DNA barcodes were used in a manner similar to that for identifying species.
In a similar way, inserted DNA barcodes are used to track sequences from a specific construct library during multiplexed sequencing (Parameswaran) or to confirm that insertional mutations originated from the viral vector and not idiopathically (Chen). DNA barcodes can be used in a similar way in almost any application that uses recombinant DNA.
DNA barcodes for gene expression studies
A form of DNA barcode technology requiring fewer manipulations is found in the nCounter® Pro Analysis System. The probes designed for nCounter Pro eliminate the need for PCR or sequencing by direct hybridization to specific target cDNAs of various gene subsets (Geiss). nCounter Pro’s panels of probes use molecular barcodes that incorporate both DNA and RNA segments.
nCounter Pro’s panels of probes use molecular barcodes that incorporate both DNA and RNA segments.
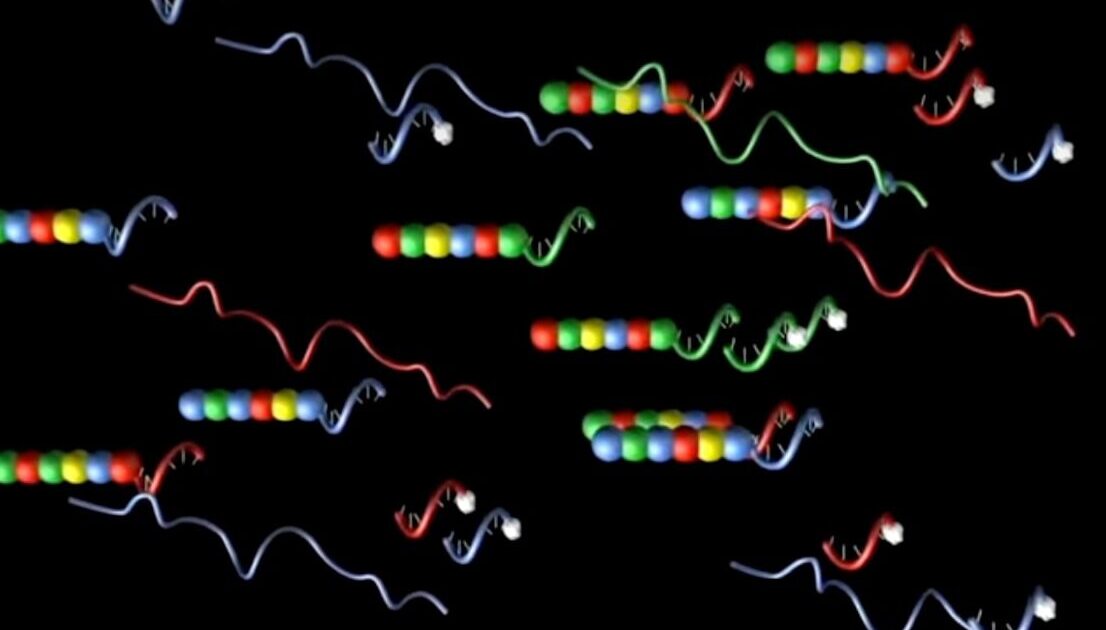
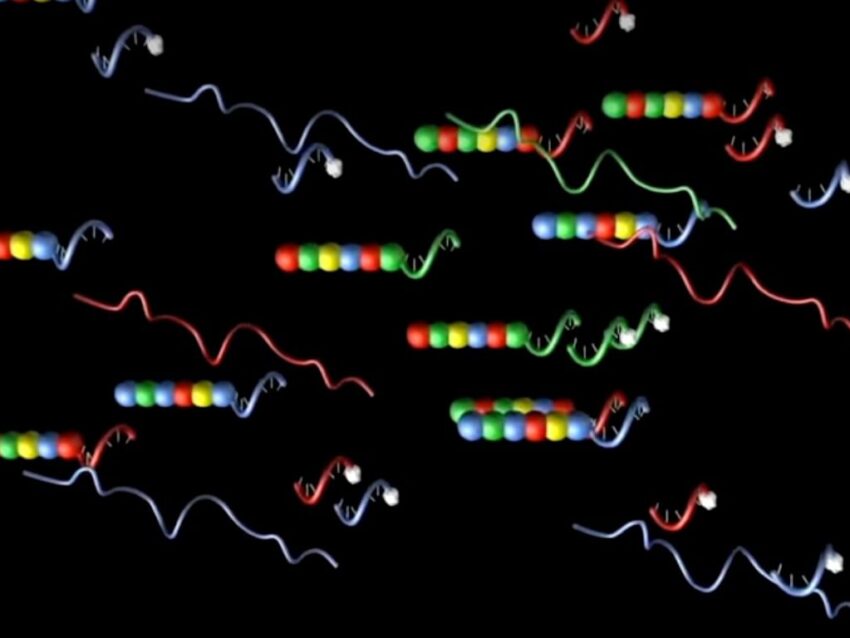
The DNA portion combines a sequence complementary to the target transcript and a sequence complementary to an in vitro transcribed RNA. Each RNA contains individual segments labeled with specific differently colored fluorophores, which are arranged in a specific linear order to create a unique code for each gene of interest. These fluorescent RNA barcodes are then annealed to the complementary single-stranded DNA portion of the probe, known as the backbone (Geiss). Ligation of repeated sequences on the 5’ end to the fluorescently labeled RNA-DNA backbone and gene-specific oligonucleotide completes the reporter half of the probe.
The second half of the two-part probe is called the capture probe, generated by fusing a gene-specific sequence adjacent to that used for the reporter probe to a series of 3’ repeats (Geiss).

After a simple hybridization protocol of the probe to target cDNA, image analysis equipment identifies each target RNA by the specific color order of the labeled reporter probe without need for any amplification or other enzymatic steps, greatly accelerating gene expression profiling.
References
Chen BR, Hale DC, Ciolek PJ, Runge KW. Generation and analysis of a barcode-tagged insertion mutant library in the fission yeast Schizosaccharomyces pombe. BMC Genomics. 2012;13:161. Published 2012 May 3. doi:10.1186/1471-2164-13-161
Geiss GK, Bumgarner RE, Birditt B, Dahl T, Dowidar N, Dunaway DL, Fell HP, Ferree S, George RD, Grogan T, James JJ, Maysuria M, Mitton JD, Oliveri P, Osborn JL, Peng T, Ratcliffe AL, Webster PJ, Davidson EH, Hood L, Dimitrov K. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat Biotechnol. 2008 Mar;26(3):317-25. doi: 10.1038/nbt1385. Epub 2008 Feb 17. PMID: 18278033.
Hebert PD, Cywinska A, Ball SL, deWaard JR. Biological identifications through DNA barcodes. Proc Biol Sci. 2003 Feb 7;270(1512):313-21. doi: 10.1098/rspb.2002.2218. PMID: 12614582; PMCID: PMC1691236.
Kress WJ, Erickson DL. DNA barcodes: genes, genomics, and bioinformatics. Proc Natl Acad Sci U S A. 2008;105(8):2761-2762. doi:10.1073/pnas.0800476105
Miller SE. DNA barcoding and the renaissance of taxonomy. Proc Natl Acad Sci U S A. 2007 Mar 20;104(12):4775-6. doi: 10.1073/pnas.0700466104. Epub 2007 Mar 15. PMID: 17363473; PMCID: PMC1829212.
Mazurkiewicz P, Tang CM, Boone C, Holden DW. Signature-tagged mutagenesis: barcoding mutants for genome-wide screens. Nat Rev Genet. 2006 Dec;7(12):929-39. doi: 10.1038/nrg1984. PMID: 17139324.
Parameswaran P, Jalili R, Tao L, et al. A pyrosequencing-tailored nucleotide barcode design unveils opportunities for large-scale sample multiplexing. Nucleic Acids Res. 2007;35(19):e130. doi:10.1093/nar/gkm760

Related Content