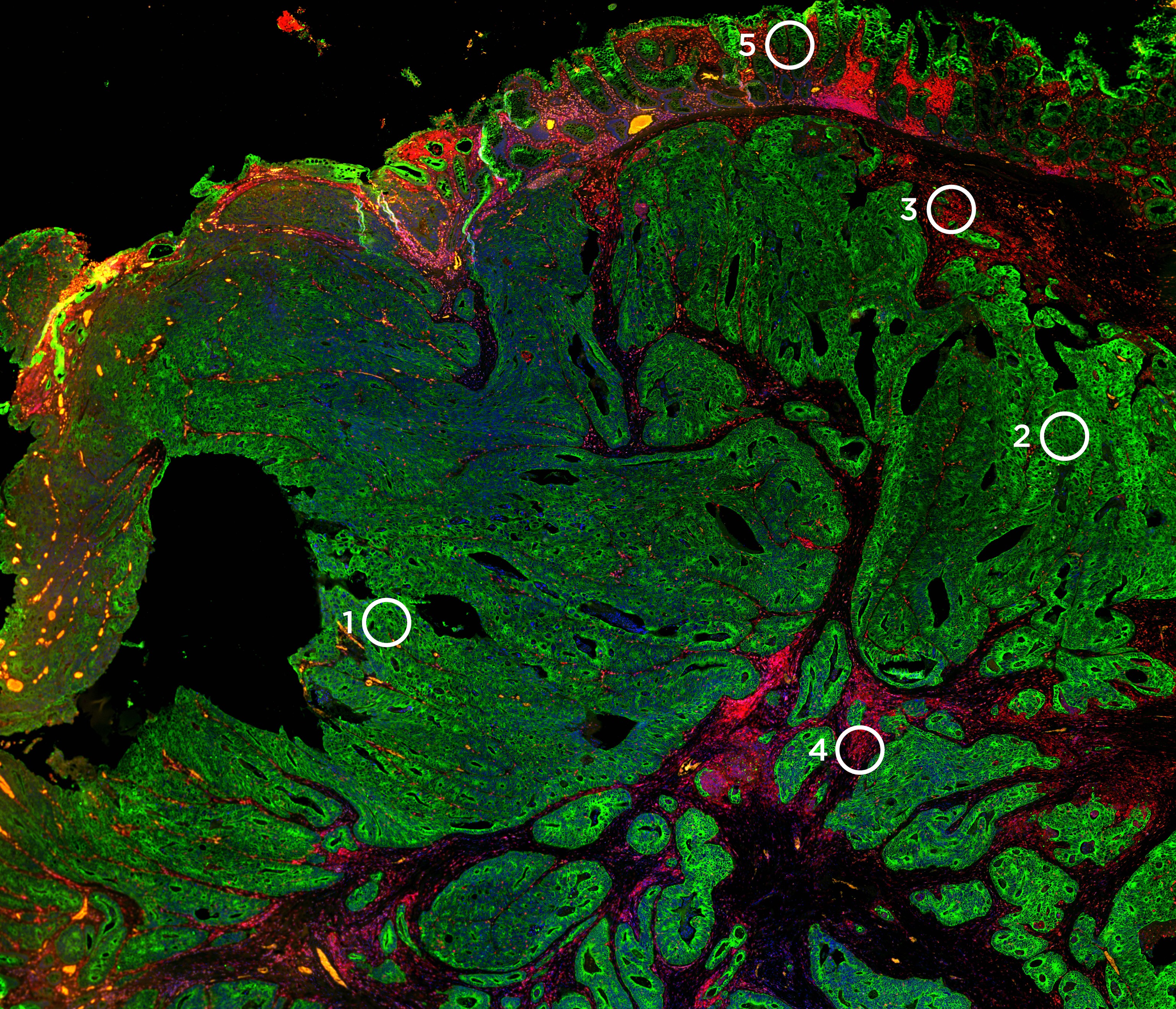
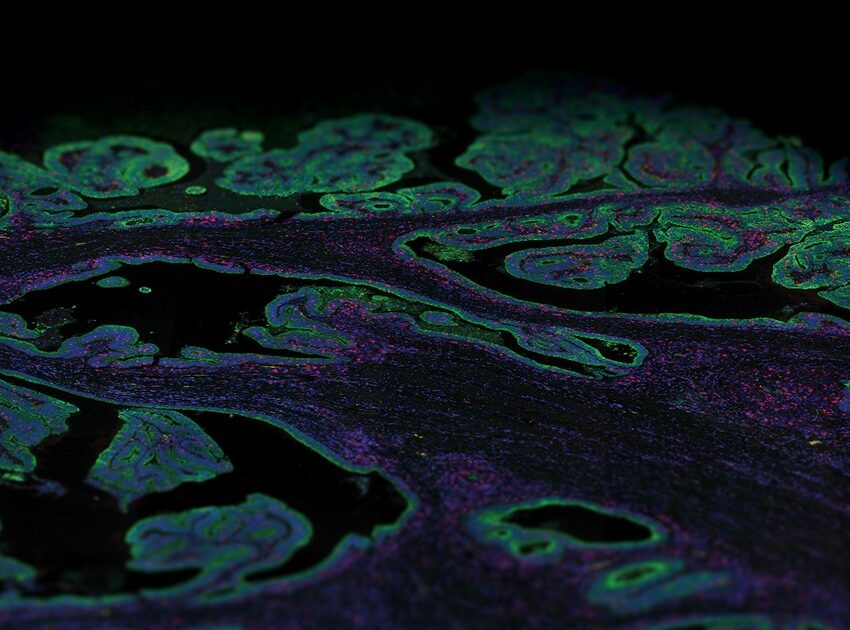
What is a morphology marker?
Understanding tissue morphology is crucial to identify and separate regions of tissue into meaningful biological groupings for image analysis of tissue sections.
This is done using morphological markers, most of which are either fluorescently linked antibodies or RNA probes that help reveal the structural architecture of tissue and define regions of interest (ROIs) to be analyzed on a tissue section. Spatial transcriptomics enables coupling high-plex expression profiling with imaging of a tissue section. Thus, morphological markers are needed to help select histological structures on the tissue section to do transcriptional profiling and to associate expression data with imaging to help identify genes associated with tissue morphology.
Morphology-driven spatial analysis
Each spatial transcriptomics method captures and stores spatial information in a variety of ways and is broadly classified into five categories: digital spatial profiling (DSP), fluorescent in situ hybridization (FISH), in situ sequencing (ISS), next-generation sequencing (NGS) with spatial barcoding, and methods that do not require a priori spatial location.

NanoSting’s GeoMx® Digital Spatial Profiler is a spatial multiomics platform that profiles the expression of RNA or proteins from a variety of sample types including formalin-fixed, paraffin-embedded (FFPE) tissue sections within user-defined ROIs. The identification of the ROI is a crucial step in the GeoMx DSP workflow. Hence, the GeoMx DSP workflow requires fluorescently labeled probes to stain for different anatomical structures, tissue compartments, and cell types. These morphology markers help guide the selection and segmentation of ROIs based on tissue structure and cell types. For instance, an ROI can be selected in immune-depleted areas or immune-enriched areas of a tumor based on appropriate staining in a tissue section.
For proteomic or transcriptomic profiling, GeoMx DSP uses oligonucleotide barcodes that are attached to fluorescent antibodies or in situ hybridization (ISH) probes, respectively, via a photocleavable linker. A pool of these barcode-labeled probes is then hybridized to RNA or protein targets on either fresh frozen or FFPE tissue sections. After fluorescent imaging, the immunofluorescent (IF) staining is used to decide where on the tissue to select different regions of interest for profiling. These ROIs are then sequentially exposed to UV light that causes the release of the oligonucleotide barcodes within the boundaries of the ROI. The released barcodes are then collected and counted with the nCounter® Analysis System or sequenced with an Illumina next-generation sequencing (NGS). The identity and quantity of the collected barcodes can then be correlated to specific RNA or proteins and digitally mapped back to the ROI on the tissue image.
Selection of morphological markers
Understanding tissue morphology is the first step in a well-designed DSP experiment.Staining of morphology markers are used to guide the selection of ROIs on the tissue, and there are several strategies to stain and image the tissue. As many spatial multi-omics platforms are a combination of fluorescent ISH techniques and some type of microscopy, the best method of visualizing morphology markers is usually by staining the tissue with primary antibodies conjugated to a fluorophore. If the antibody of choice is not labeled with the preferred fluorophore, there are several kits available that fluorescently label antibodies through direct conjugation. The best type of antibodies for a spatial biology experiment are monoclonal antibodies due to the ideal signal-to-noise ratio, but polyclonal antibodies have also been shown to work on GeoMx DSP.
There are numerous morphology markers available from NanoString developed as morphology kits that contain common tissue markers. Further, there is a list of previously validated morphology markers for GeoMx assays available. Another method for staining the tissue for visualization is using ISH probes as morphology markers; a wide selection of RNA targets are available as ISH probes. RNAscope, a commercially available method for ISH, has been validated to work with the GeoMx DSP workflow.
The GeoMx DSP control software contains an image overlay tool that allows for the placement of ROIs using serial sections of tissue prepared using IF or hematoxylin & eosin (H&E) staining. This allows for tissue staining and ROI selection to be done separately or remotely by a pathologist at a different location than the GeoMx instrument.